Diagnostic and prognostic models are typically evaluated with measures of accuracy that do not address clinical consequences. Decision-analytic techniques allow assessment of clinical outcomes but often require collection of additional information may be cumbersome to apply to models that yield a continuous result. Decision curve analysis is a method for evaluating and comparing prediction models that incorporates clinical consequences, requires only the data set on which the models are tested, and can be applied to models that have either continuous or dichotomous results. The dca function performs decision curve analysis for binary outcomes. Review the DCA Vignette for a detailed walk-through of various applications. Also, see www.decisioncurveanalysis.org for more information.
Arguments
- formula
a formula with the outcome on the LHS and a sum of markers/covariates to test on the RHS
- data
a data frame containing the variables in
formula=.- thresholds
vector of threshold probabilities between 0 and 1. Default is
seq(0, 0.99, by = 0.01). Thresholds at zero are replaced with 10e-10.- label
named list of variable labels, e.g.
list(age = "Age, years")- harm
named list of harms associated with a test. Default is
NULL- as_probability
character vector including names of variables that will be converted to a probability. Details below.
- time
if outcome is survival,
time=specifies the time the assessment is made- prevalence
When
NULL, the prevalence is estimated fromdata=. If the data passed is a case-control set, the population prevalence may be set with this argument.
Value
List including net benefit of each variable
as_probability argument
While the as_probability= argument can be used to convert a marker to the
probability scale, use the argument only when the consequences are fully
understood. For example, when the outcome is binary, logistic regression
is used to convert the marker to a probability. The logistic regression
model assumes linearity on the log-odds scale and can induce
miscalibration when this assumption is not true. Miscalibration in a
model will adversely affect performance on decision curve
analysis. Similarly, when the outcome is time-to-event, Cox Proportional
Hazards regression is used to convert the marker to a probability.
The Cox model also has a linearity assumption and additionally assumes
proportional hazards over the follow-up period. When these assumptions
are violated, important miscalibration may occur.
Instead of using the as_probability= argument, it is suggested to perform
the regression modeling outside of the dca() function utilizing methods,
such as non-linear modeling, as appropriate.
Examples
# calculate DCA with binary endpoint
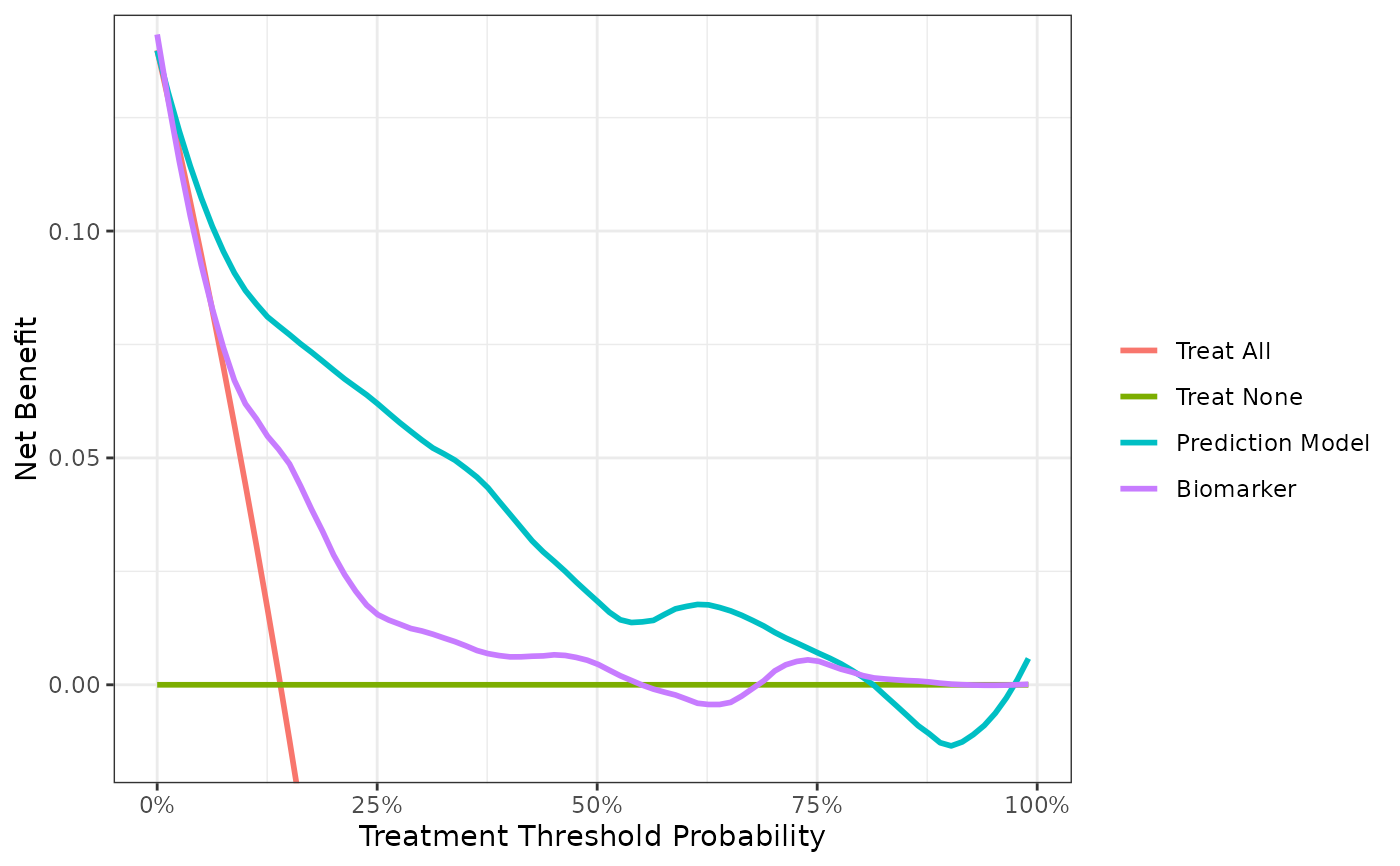
dca(cancer ~ cancerpredmarker + marker,
data = df_binary,
as_probability = "marker",
label = list(cancerpredmarker = "Prediction Model", marker = "Biomarker")) %>%
# plot DCA curves with ggplot
plot(smooth = TRUE) +
# add ggplot formatting
ggplot2::labs(x = "Treatment Threshold Probability")
 # calculate DCA with time to event endpoint
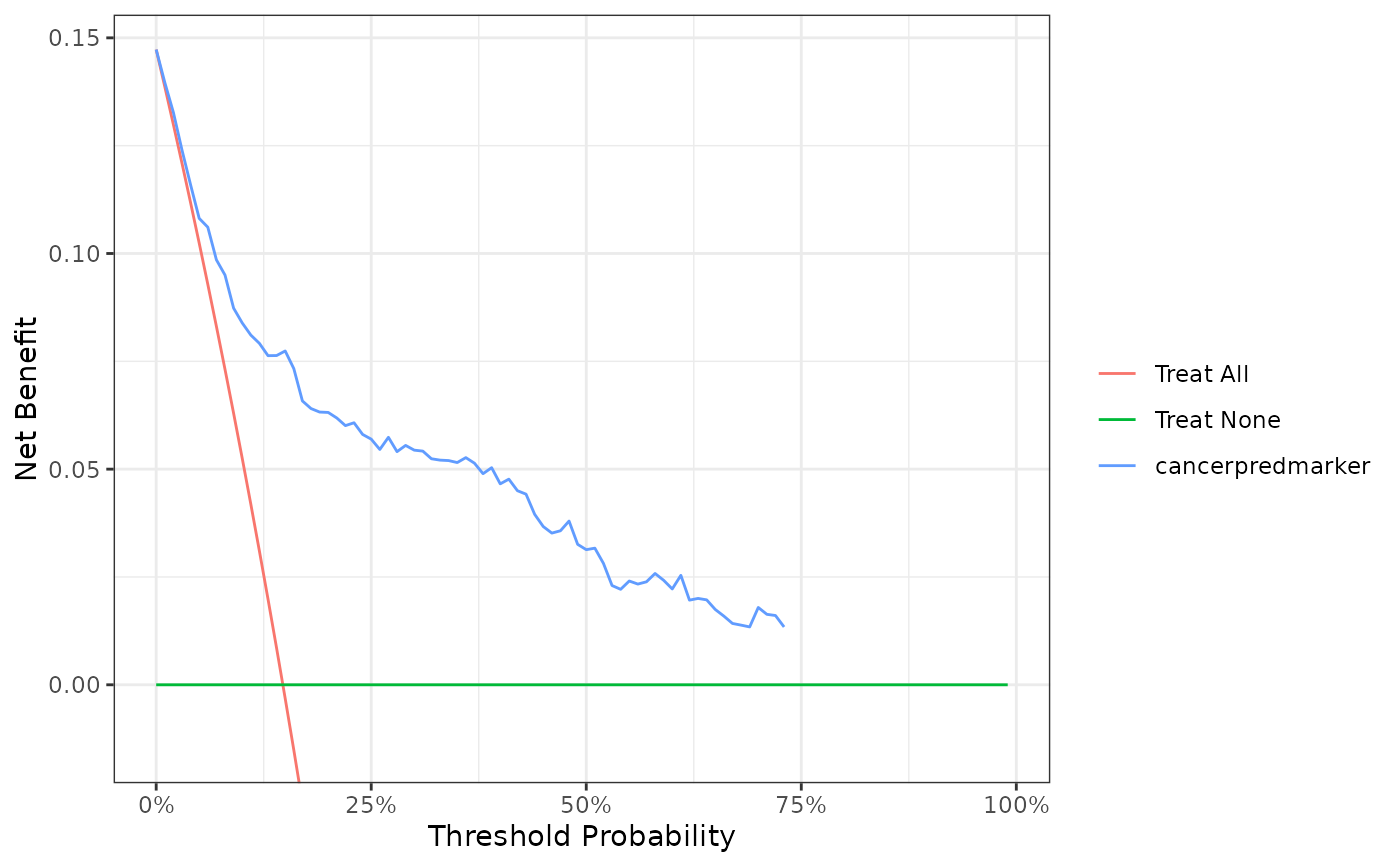
dca(Surv(ttcancer, cancer) ~ cancerpredmarker, data = df_surv, time = 1)
#> Printing with `plot(x, type = 'net_benefit', smooth = FALSE, show_ggplot_code = FALSE)`
# calculate DCA with time to event endpoint
dca(Surv(ttcancer, cancer) ~ cancerpredmarker, data = df_surv, time = 1)
#> Printing with `plot(x, type = 'net_benefit', smooth = FALSE, show_ggplot_code = FALSE)`